Tanglegrams are a visual method to compare two phylogenetic trees with the same set of tip labels. This can be useful for comparing trees produced by different methods on the same alignment, or on different alignments of the sample set. Tanglegrams work by connecting the matching tips of the trees, then rotating subtrees to minimise the number of crossings. The algorithm was published in 2011, and continues to be used in a range of publications (for example genomic epidemiology).
I recently ran into an issue using tanglegrams which I’d like to document here. I constructed phylogenies from the protein alignment of all the alleles of PspA identified in a carriage collection of Streptococcus pneumoniae using RAxML (a maximum likelihood method) and mrbayes (a Bayesian method which produces a posterior of trees). For the latter, one can use the R package treescape to find the median tree in the posterior distribution. I wanted to compare the shape of these two trees to see if there were any obvious areas of uncertainty in the topology (a perhaps more quantitative way to do this might be by looking at bootstrap values). I made a tanglegram, from which my initial interpretation was trees were very similar:
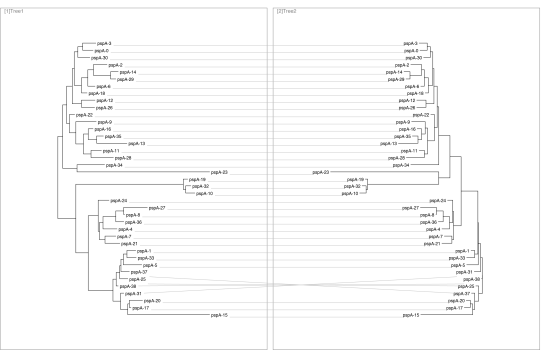
pspA allele trees. Tanglegram between maximum likelihood tree and median from the Bayesian posterior.
There is only one clade of swaps (near the bottom), so the trees must be similar? Not necessarily. A game I like to play is manually rotating subtrees to make metadata cluster. Though the topology remains identical, visually the clustering of metadata can change drastically. Even though we know vertical distance is meaningless in a phylogeny, the way this is usually visually displayed can lead us to draw erroneous conclusions.
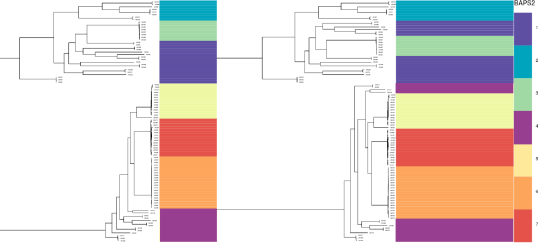
Rotating subtrees. Left: nice well defined clades clustering together. Right: same tree topology and labels, but the clades no longer form single clusters. Clusters 1 and 4 are polyphyletic, and rotating subtrees can break up or combine these vertically.
Let’s look at the tanglegram above again in more detail:

Misleading tanglegram? This emphasises topology changes in more recent parts of the phylogeny (blue) and may be misleading about similarity of ancestral branches (red).
The swap highlighted in blue is the only event where tip lines cross. This correctly leads us to identify that pspA-31 and pspA-37 are in slightly different places in this part of the phylogeny. However look in the red box. Those these strains totally align, the placement of them within the phylogeny is quite different (red arrows)! In this case I don’t think the line crossings by the tanglegram have identified what I wanted to see here, which is that the ancestral topology of these two trees is different.
While I am not claiming tanglegrams are not useful, without careful attention they may be misleading. A good alternative to tanglegrams, in my opinion, is phylo.io. This allows you to dynamically compare two tree topologies, and see how each branch flip affects the arrangement rather than providing just one static comparison. If you want to try this out on the trees above the two newick files are attached at the end of this post to copy in. Finally, I would also note that there are various metrics which can provide a quantitative comparison between (many) trees, my favourite of which is the Kendall-Colijin metric implemented in treescape.
Update: Michelle Kendall (an author of both the KC metric and treescape) pointed out on twitter that they have added the function plotTreeDiff() as another way to look at topology changes. I gave this a go:

plotTreeDiff result. The two trees, with tips coloured based on how many differences in recent common ancestors there are with other tips.
From the manual:
‘A plot of the two trees side by side. Tips are coloured in the following way:
- if each ancestor of a tip in tree 1 occurs in tree 2 with the same partition of tip descendants, then the tip is coloured grey (or supplied “baseCol”)
- if not, the tip gets coloured pale orange to red on a scale according to how many differences there are amongst its most recent common ancestors with other tips. The colour spectrum can be changed according to preference.’
You can use tipDiff() to see the numeric results direcly.
Trees:
((((((pspA-15:0.7542002,(pspA-20:0.1224276,pspA-17:0.01459105):0.0136343):0.06696656,pspA-31:0.02844112):0.07110159,(pspA-38:0.0188037,((((pspA-1:0.07786275,pspA-33:0.08346416):0.03494853,pspA-5:0.1691264):0.02173634,pspA-37:0.0787912):0.009598778,pspA-25:0.07136204):0.03492139):0.02958377):0.2273015,(((pspA-7:0.09993183,pspA-21:0.1386982):0.05063578,(pspA-4:0.1303186,((pspA-27:0.2173186,pspA-8:0.006066717):0.07263402,pspA-36:0.07059364):0.119125):0.03385501):0.01120214,pspA-24:0.09906705):0.09046543):0.1139506,(pspA-19:0.04005775,(pspA-32:0.04931893,pspA-10:0.08485811):0.02024913):0.9919588):0.167318775,((((((pspA-3:0.1203388,pspA-0:0.09452877):0.04488296,pspA-30:0.1090477):0.03244832,((pspA-6:0.06727758,((pspA-29:0.04877083,pspA-14:0.07255919):0.1581595,pspA-2:0.1249573):0.0518224):0.05925152,pspA-18:0.05166722):0.06424128):0.01986727,(pspA-12:0.1279069,pspA-26:0.1232956):0.08339421):0.05060685,(pspA-22:0.04905987,(pspA-9:0.1901771,((pspA-16:0.04409937,(pspA-35:0.06477806,pspA-13:0.2830619):0.07727303):0.04340311,(pspA-11:0.09526805,pspA-28:0.2083244):0.07013518):0.07074651):0.05979277):0.02315286):0.03699984,(pspA-34:0.2528056,pspA-23:1.222536):0.1333402):0.044963025);
(((pspA-5:0.16557530729568833983,((pspA-1:0.04756995109125589788,pspA-33:0.10915366994507740006):0.04785917480955029918,(((pspA-7:0.10230120656769127463,pspA-21:0.14095901783884545733):0.02506804670603787408,(((pspA-36:0.06861655800623776835,(pspA-8:0.00442478384355664365,pspA-27:0.22142270295956195669):0.09444585738499523819):0.08879457852006654439,pspA-4:0.10604010012498631121):0.04672183194168183507,pspA-24:0.07323682699184425049):0.01199257548493264970):0.10529470578563034089,((pspA-34:0.38604601286039375019,(((pspA-12:0.12639557827350531016,pspA-26:0.12595340725708714658):0.11169806126353225284,((pspA-18:0.04075294523735660535,(pspA-6:0.05097275166368370192,((pspA-14:0.08245346709560481824,pspA-29:0.05470945144269170890):0.16281232925454419691,pspA-2:0.15070678273765572563):0.03938700493100827371):0.05203089575251463456):0.06302461915576125506,(pspA-30:0.09148296281857748458,(pspA-0:0.10463631384718696804,pspA-3:0.11382571714494262027):0.04249311451680449353):0.03340348219787352829):0.01489076329975322355):0.04853549722852742304,(pspA-22:0.04696709784392495701,((pspA-11:0.07535697579734597362,pspA-28:0.25243204747960418244):0.04403388452890973775,(pspA-9:0.18996034668835029557,(pspA-16:0.06144862376464669401,(pspA-13:0.27683177216876692084,pspA-35:0.07992819540786025301):0.05138818922505713344):0.04831192529255548540):0.03498701951880421601):0.09272730151798472265):0.04073871346767179297):0.01853343949682505903):0.15648667630311274834,((pspA-19:0.03232176246079743881,(pspA-10:0.09060060658061755423,pspA-32:0.05569822158278419505):0.02219077947076726620):0.99299057857468620014,pspA-23:1.51276427797469659176):0.15413589497708032883):0.15939898234062221949):0.23660076759918716172):0.00461861925303937333):0.03002044039766965655,pspA-31:0.09022645321942507346):0.02410560069003805581,(((pspA-15:0.72495676640385564582,(pspA-17:0.08537518563518509129,pspA-20:0.05450107613074649943):0.03312225101520550885):0.02823410020886808411,pspA-37:0.04303607734486421255):0.05520151224297501630,pspA-25:0.04043049259731745781):0.05152615294739129603,pspA-38:0.00917388377063698898):0.0;



